Most nervous systems are largely bilaterally symmetric on a structural level, but are usually functionally lateralized[1]. How left/right asymmetry is developmentally controlled is poorly understood, in part because there are few molecular correlates to functional asymmetries in any nervous system. The C.elegans nervous system displays a number of striking asymmetries in its nervous system [1]. We have focused much of our efforts on the two bilaterally symmetric gustatory neurons ASE(Left) and ASE(Right) as an entry point into studying how asymmetry develops.
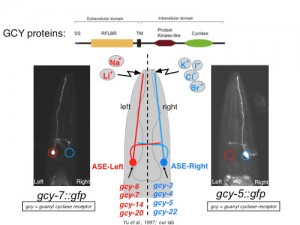
ASE(L) and ASE(R) express in a stereotyped, left/right asymmetric manner many members of a receptor-type guanylyl cyclase (rGCY) family that we found to be required for left/right asymmetric taste perception (Figure 1)[2, 3]. The ability to easily visualize this functional asymmetry through gfp reporters that tag therGCY-encoding gcy loci has therefore provided us with a unique opportunity to ask how asymmetry is developmentally programmed. Using three different approaches, we have provided the following insights:
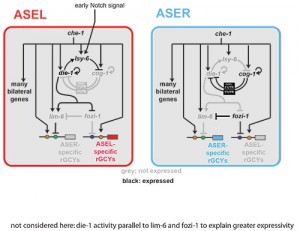
(i) We have dissected cis-regulatory regions of the gcy genes to probe the regulatory logic of their left/right asymmetric restriction, revealing that they are under direct control of the bilaterally expressed terminal selector CHE-1, but that additional cis-regulatory motifs promote or inhibit the activity of CHE-1 in either ASE(L) or ASE(R)(Figure 2)[4].
(ii) Through genetic screens we have elucidated a complex network of regulatory RNAs, transcription factors and chromatin modifiers, many under direct control of the terminal selector CHE-1, that regulate asymmetric gcy expression (Figure 2)[4-16].
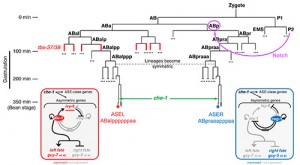
(iii) Early embryonic blastomere manipulations revealed that the initial trigger for ASE(L/R) asymmetry lies in a Notch signal sent in the early embryo, long before the ASE neurons are born (Figure 3)[7]. Via intermediary T-box transcription factors, this signal induces a decompaction of chromatin at the lsy-6 miRNA locus throughout embryonic development in the ABa lineage. lsy-6 expression is then amplified in the ASE(L) neuron through the terminal selector che-1 and the bistable feedback loop shown in Figure 3 to trigger postmitotic, asymmetric differentiation of the ASE(L/R) neurons[17].
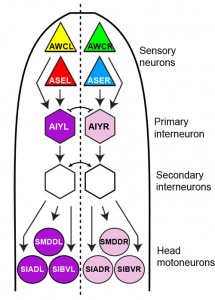
We have also found that asymmetry in the C.elegans nervous system extends beyond chemosensory asymmetry. We have identified left/right asymmetric gene expression programs in several interneurons and motoneurons and have shown that they are induced by a novel, mesodermally derived Notch signal[18](Figure 4). These findings reveal a previously underappreciated complexity in the neuronal circuitry of the worm.
A key factor in ASE left/right asymmetry control is the ASEL-specific miRNA, lsy-6, which we had discovered previously [19]. Aside from placing a miRNA in a complex gene regulatory network (Figure 3), we have used the lsy-6 interaction with its genetically defined target cog-1, to probe concepts of miRNA/target interactions, thereby challenging some commonly accepted rules[20, <21]. As we detailed in an essay in Science[22], this and other work reveals striking conceptual similarities between miRNA and transcription factor action.